Building a Reproducible Classical–Quantum ML Platform for Molecular Prediction
Source: Dev.to
Source: Dev.to – Building a Reproducible Classical‑Quantum ML Platform for Molecular Prediction
Why Most Quantum ML Demos Fall Short
Quantum machine learning is often demonstrated through small, highly controlled examples: tiny datasets, shallow circuits, and weak baselines. While useful for education, these setups rarely convince practitioners.
Common Issues
- Datasets too small – they don’t stress model capacity.
- Weak classical baselines – comparisons are unfair.
- High variance – stochastic quantum training leads to noisy results.
- Lack of reproducibility – little attention to reproducible pipelines or deployment.
Instead of building “a quantum model,” this project treats quantum ML the same way we treat any new modeling family in applied ML:
- A shared, reproducible experimental platform.
- Strong, realistic baselines.
- Rigorous evaluation.
- Production‑ready considerations.
End‑to‑End Platform Overview
This project is not a single model but a config‑driven research and engineering framework for molecular property prediction. It supports three model families under a unified pipeline:
- Classical graph neural networks (GNN, GAT, MPNN + GRU + Set2Set)
- Quantum models (variational circuits and quantum kernels)
- Hybrid classical‑quantum models with learned fusion
All models share the same:
- Data ingestion
- Featurisation
- Batching strategy
- Training loop
- Evaluation metrics
This design ensures that differences in performance can be attributed to architecture, not hidden changes in preprocessing or optimisation.

From SMILES to Molecular Graphs
Molecules are supplied as SMILES strings—compact but not directly aligned with the relational nature of chemical interactions. To expose this structure, each SMILES string is transformed into a molecular graph using RDKit.
| Graph Component | Description |
|---|---|
| Nodes | Atoms |
| Edges | Bonds (directed to enable message‑passing) |
Atom Features
Chemically meaningful properties encoded for each atom:
- Atom type (element symbol)
- Atomic degree (number of directly bonded neighbors)
- Hybridisation (sp, sp², sp³, …)
- Formal charge
- Aromaticity flag
Bond Features
Properties captured for each bond:
- Bond type (single, double, triple, aromatic)
- Conjugation flag
- Ring membership flag
These features provide the foundation for downstream graph‑based models that learn from the true relational structure of molecules.
Why Geometry Matters (Especially for QM9)
Many molecular properties depend not only on connectivity but also on three‑dimensional geometry. This is particularly true for the electric dipole moment ( \mu ), which is governed by the spatial distribution of partial charges.
The platform can optionally augment molecular graphs with 3‑D features:
- Add hydrogen atoms.
- Generate a conformer using RDKit’s ETKDG algorithm.
- Relax the structure via MMFF optimisation.
- Each atom receives Cartesian coordinates and its distance to the molecular centroid.
- Edges are augmented with bond‑length information.
Note: The same featurisation pipeline feeds all model families, isolating architectural effects rather than input differences.
Scaling Graph Learning in JAX
Graph neural networks introduce a practical challenge: molecules have variable size, which clashes with JAX’s preference for static tensor shapes.
The platform introduces a BatchedGraph abstraction:
- Padding – Node and edge features are padded to the batch‑level maxima.
- Masks – Boolean masks track real vs. padded entries.
- Mask‑aware ops – All aggregation and pooling operations respect the masks.
Benefits
- ✅ True mini‑batch training
- ✅ JIT‑compiled forward passes
- ✅ Stable gradient statistics
- ✅ Efficient hyper‑parameter sweeps
In practice, this approach yields roughly an order‑of‑magnitude speed‑up over naïve per‑graph execution.
Three Model Families Under One Roof
Classical Baselines That Actually Matter
A rigorous evaluation of quantum or hybrid models requires strong classical baselines. The platform provides:
- Message‑Passing GNN – a standard graph‑convolutional baseline.
- Graph Attention Network (GAT) – edge‑aware attention for richer relational modeling.
- MPNN with GRU updates & Set2Set pooling – combines recurrent updates with a powerful read‑out.
All three models share the same training‑ and evaluation‑pipeline, guaranteeing a fair comparison.
Quantum Models Beyond Toy Circuits
A legacy 4‑qubit variational circuit is included as a reference point (lower bound on quantum capacity). More expressive quantum models are then explored:
| Model | Key Features |
|---|---|
| 8‑qubit variational circuits | Larger Hilbert space, deeper expressivity. |
| Data‑reuploading feature maps | Re‑inject classical data at multiple layers to boost representational power. |
| Hardware‑aware circuit design | Tailored to the connectivity and native gate set of the target quantum processor. |
| Quantum kernel methods | Non‑variational alternative that leverages quantum‑enhanced similarity measures. |
Noise, shot statistics, and gradient behaviour are treated as first‑class concerns rather than ignored artifacts.
Hybrid Classical‑Quantum Models Done Properly
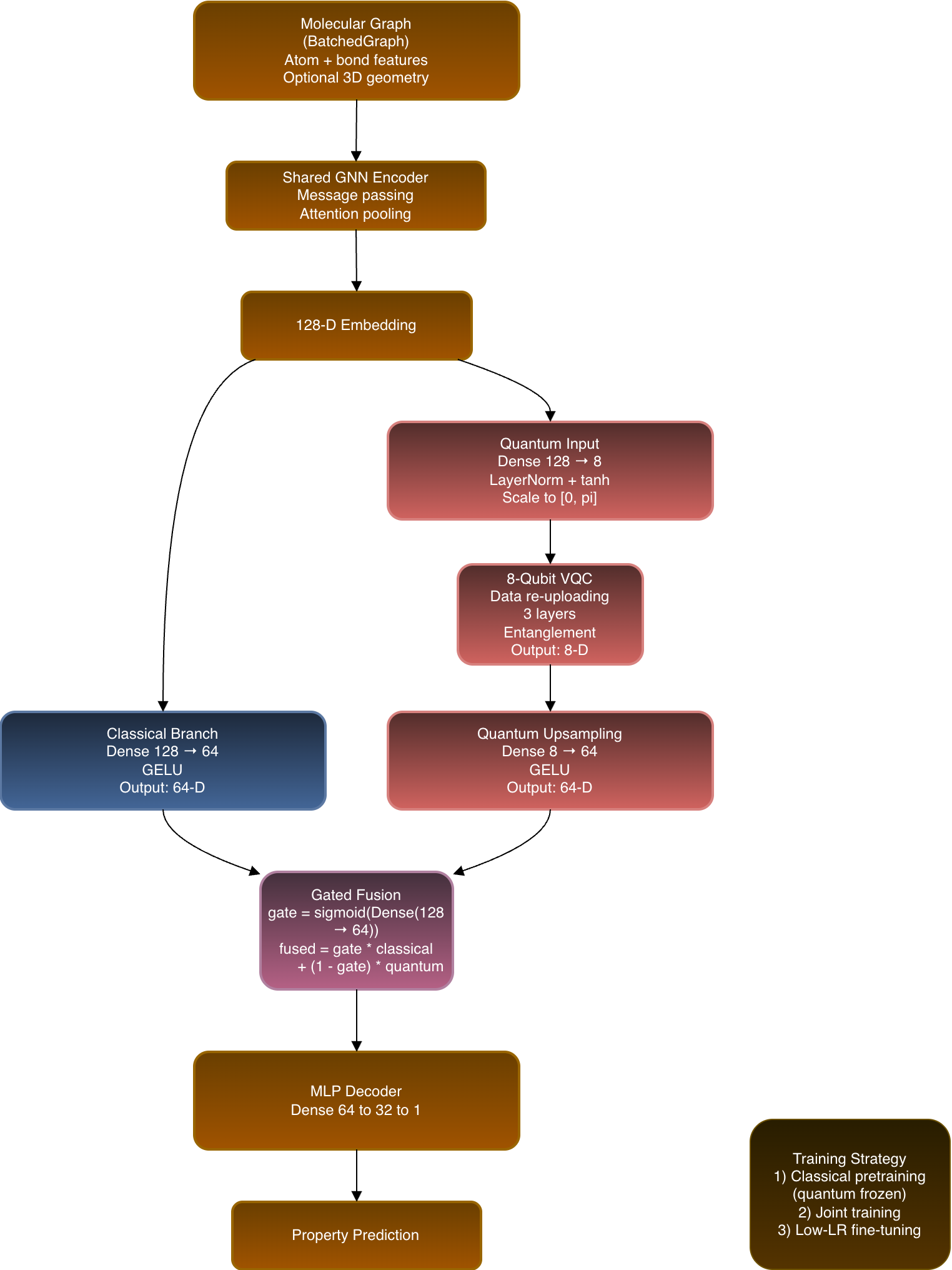
Naïve hybrids often concatenate a tiny quantum output with a large classical embedding, causing the quantum signal to be drowned out. The Hybrid V2 architecture enforces dimension‑matched fusion:
- Shared GNN encoder → 128‑dimensional node/graph embedding.
- Classical branch → projects the 128‑D embedding to 64 D.
- Quantum branch
- Compresses the 128‑D embedding into a set of qubits.
- Processes it with a variational quantum circuit (VQC).
- Upsamples the quantum output back to 64 D.
- Learned gating layer → combines the 64‑D classical and quantum representations per input.
Training Schedule
| Stage | Description |
|---|---|
| 1️⃣ Classical pre‑training | Train the GNN and classical branch alone until convergence. |
| 2️⃣ Joint training | Enable the quantum branch and train the full hybrid end‑to‑end. |
| 3️⃣ Low‑learning‑rate fine‑tuning | Refine all parameters with a reduced learning rate to stabilize quantum contributions. |
This staged approach lets quantum features contribute conditionally, rather than being assumed to improve performance automatically.
Illustration
(The original image link was incomplete, so the illustration has been omitted.)
Training, Evaluation, and Scientific Rigor
Experiments are defined through configuration files and hashed for reproducibility. Random seeds are controlled across Python, NumPy, and JAX.
Evaluation includes
- Murcko scaffold splits – realistic generalisation across chemical space.
- Systematic ablations of architecture and circuit parameters.
- Uncertainty estimation via Monte‑Carlo dropout and quantum shot noise.
- Diagnostics for barren plateaus and overall trainability.
- Surrogatability tests to probe how well classical models can mimic quantum outputs.
The goal is not to chase headline numbers but to gain a deeper understanding of model behaviour.
From Research Code to Production Inference
Trained models are managed via a lightweight model registry that handles configuration, checkpoint loading, and caching.
A FastAPI service exposes:
- Single‑molecule inference
- Batch inference
- Optional uncertainty estimates
From the client’s perspective, classical, quantum, and hybrid models share the same interface, closing the gap between experimentation and real‑world use.
Code and Reproducibility
The full implementation—including configuration files, data loaders, model architectures, training pipelines, evaluation utilities, and the FastAPI inference service—is available in the GitHub repository:
GitHub repository:
https://github.com/your-username/your-repo
Final Thoughts
Rather than asking whether quantum models “win”, this platform asks a more useful question: when, where, and how do they contribute?
Answering that responsibly requires shared infrastructure, strong baselines, and measurable uncertainty. This project aims to provide exactly that.