임상 데이터 분석 자동화: 병원 데이터 내보내기에서 논문 초안까지 파이프라인
Source: Dev.to
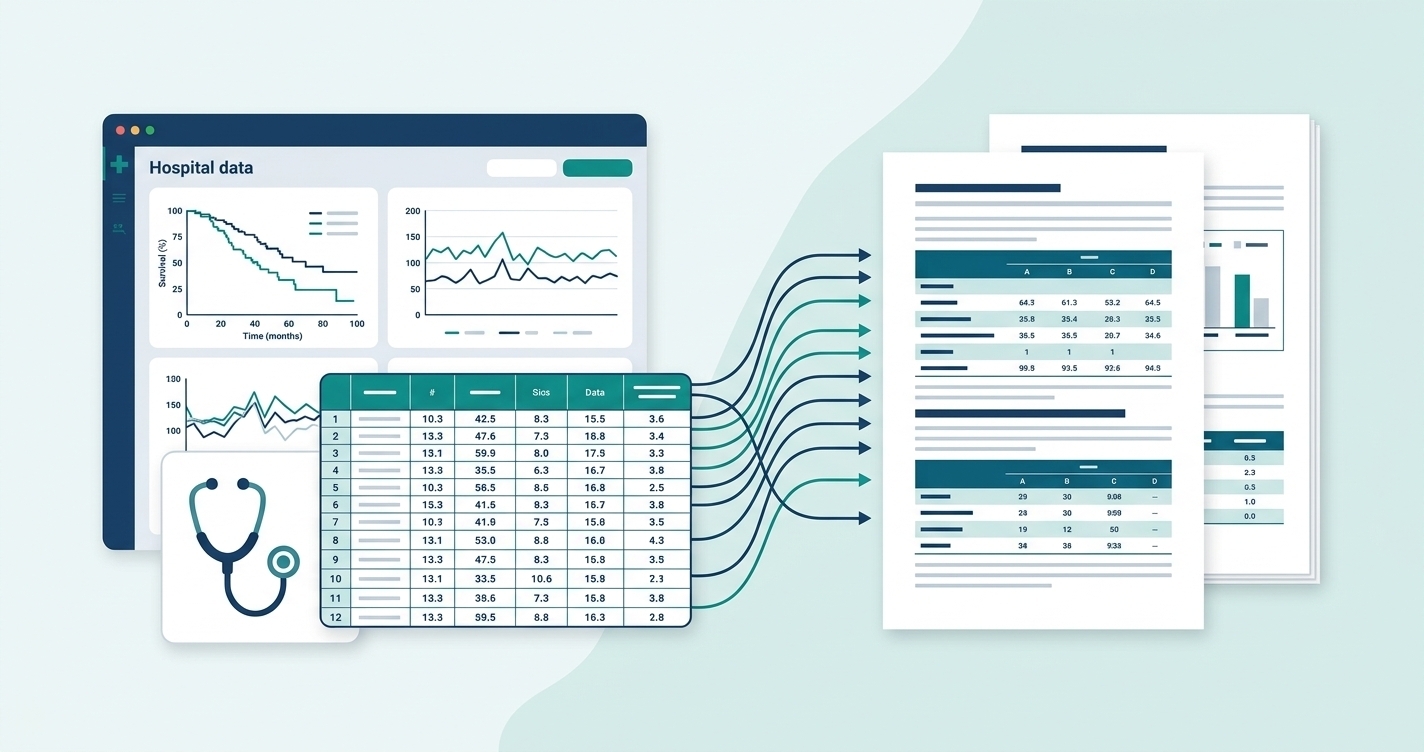
The input problem
A typical clinical data export looks like this:
PatientID | Age | Sex | HbA1c | SBP | DBP | eGFR | Dx | AdmDate | DisDate | Status
001 | 67 | M | 8.2 | 145 | 92 | | T2DM | 2024-01-15 | 01/25/2024 | alive
002 | 54 | F | | 128 | 78 | 85 | 2型糖尿病 | 20240203 | 2024-02-10 |
003 | -5 | M | 7.1 | 300 | 85 | 92 | type 2 DM | 2024-03-01 | 2024-03-08 | deadNotice: three different date formats in the same column, the same diagnosis coded three different ways, an obviously wrong age, a systolic BP that is likely a data‑entry error, missing values that could mean “not tested” or “not recorded,” and mixed languages. This variability is typical for clinical exports.
The analysis pipeline
Raw export (CSV/XLSX)
│
├─ Structure detection
│ └─ row = patient? visit? wide? long?
│
├─ Data cleaning
│ ├─ Date format standardization
│ ├─ Coding unification ("T2DM" = "2型糖尿病" = "type 2 DM")
│ ├─ Outlier flagging (SBP=300, Age=-5)
│ └─ Missing value classification (not tested vs not recorded)
│
├─ Variable typing
│ ├─ Continuous (age, HbA1c, eGFR)
│ ├─ Categorical (sex, diagnosis, comorbidities)
│ └─ Time‑to‑event (survival time + censoring status)
│
├─ Statistical analysis (Python execution)
│ ├─ Baseline table with per‑variable test selection
│ ├─ Regression (logistic / Cox / linear / Poisson)
│ ├─ Survival analysis (KM + log‑rank)
│ └─ Diagnostic evaluation (ROC + AUC)
│
└─ Output generation
├─ Formatted tables (baseline, regression results)
├─ Figures (KM curves, ROC curves, forest plots)
└─ Manuscript sections (methods + results)Key technical decisions
- Python execution, not LLM computation. Statistics must be verifiable. The LLM writes the interpretation;
scipy,statsmodels, andlifelinescompute the numbers. - Clinical variable lookup. Recognizing “SBP” as systolic blood pressure enables domain‑aware outlier detection (e.g., flag 300 mmHg as likely error) rather than relying solely on statistical outlier methods.
- Assumption checking. Every statistical test includes prerequisite verification—normality for parametric tests, events‑per‑variable for logistic regression, proportional hazards for Cox models. Running analysis without these checks is the #1 reason clinical papers get sent back by reviewers.
The baseline table problem
Generating Table 1 (baseline characteristics) sounds simple but requires per‑variable logic:
for variable in dataset:
if is_categorical(variable):
# n (%), chi‑square or Fisher's exact
elif is_normal(variable):
# mean ± SD, t‑test or ANOVA
elif is_skewed(variable):
# median (IQR), Mann‑Whitney or Kruskal‑WallisThe tricky part is automating the normality decision and handling edge cases (e.g., small cell counts triggering Fisher’s exact test instead of chi‑square).
Stack
- Next.js + Vercel
- Claude API for text generation
- Python chain for statistical computation
- Export formats: PDF / DOCX / LaTeX / ZIP
- 7 output languages
What I’m still figuring out
- Better heuristics for distinguishing “not tested” vs “not recorded” missing values
- Automated detection of wide vs long format in longitudinal datasets
- Handling mixed‑language clinical notes in the same dataset
If you’ve worked on similar problems—clinical data pipelines, automated statistical analysis, or structured document generation from data—I’d love to compare notes.